The Computational Unit, headed by Dr. Georg Gerber, collaborates with investigators on experimental design and advanced computational analyses for microbiome studies. Our focus is on innovative projects, particularly those that involve longitudinal studies of the microbiome in the context of human disease.
Examples of collaborative projects completed or ongoing in the Unit include:
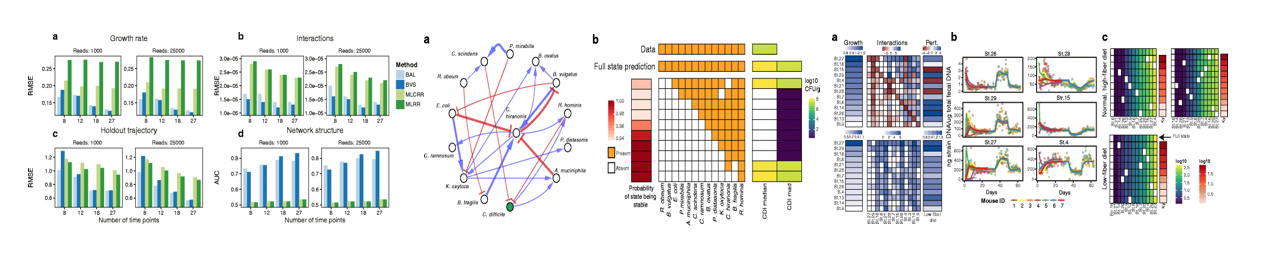
- Predicting microbial dynamics and interactions from time-series data, with applications to developing bacteriotherapies for Clostridium difficile infection and inflammatory bowel disease.
- Determining microbial genes that enhance fitness in the gut environment from functional metagenomics data.
- Modeling the dynamic effects of antibiotics or infection on the microbiota.
- Characterizing longitudinal differences in the microbiota in mouse models of immune dysfunction versus normal controls.
- Identifying microbial communities in the gut that differentiate human subjects with multiple sclerosis or food allergies from controls.
Investigators interested in collaborating are encouraging to contact us at early stages of a project, as we have found that thoughtful planning and solid experimental designs are essential to the success of microbiome projects.
Hardware resources available in the Unit include dedicated and shared high-speed storage and compute nodes on the Partners ERISOne Linux Cluster, which has an aggregate of over 350 compute nodes, 4500 CPU cores and a total of 25TB RAM memory and 2PB’s of storage in addition to specialized parallel processing resources. Software used by the Unit includes customized bioinformatics pipelines for 16S rRNA and metagenomics analysis, as well as advanced machine learning algorithms for analyzing microbiome dynamics developed by the Gerber lab.
Case Studies:
MDSINE: Microbial Dynamical Systems Inference Engine
Pathogen Genomic Surveillance Program
To inquire further please fill out our online request form or contact us at microbiome@bwh.harvard.edu.
